import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from scipy.stats import ttest_ind
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, roc_auc_score, roc_curve
from sklearn.preprocessing import StandardScaler
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import DenseAbout Data
According to the provided information, the medical dataset classifies either heart attack or none. The gender column in the data is normalized: the male is set to 1 and the female to 0. The glucose column is set to 1 if it is > 120; otherwise, 0. As for the output, positive is set to 1 and negative to 0.
The dataset has 9 column:
- Age: The patient’s age
- Gender: Biological sex of the patient (The male is set to 1 and the female to 0)
- Heart Rate: The number of heartbeats per minute
- Systolic Blood Pressure: The pressure in arteries when the heart contracts
- Diastolic Blood Pressure: The pressure in arteries between heartbeats
- Blood Sugar: The patient’s blood glucose level
- Ck-mb: A cardiac enzyme released during heart muscle damage
- 심장 근육에 존재하는 효소로, 심장 근육이 손상되면 방출된다.
- Troponin:A highly specific protein biomarker for heart muscle injury
- 심장 근육 수축에 관여하는 단백질
- Result: The outcome label indicating whether or not the patient experienced a heart attack
Import
Data
df = pd.read_csv("../../../../delete/Medicaldataset.csv")
df.head()| Age | Gender | Heart rate | Systolic blood pressure | Diastolic blood pressure | Blood sugar | CK-MB | Troponin | Result | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 64 | 1 | 66 | 160 | 83 | 160.0 | 1.80 | 0.012 | negative |
| 1 | 21 | 1 | 94 | 98 | 46 | 296.0 | 6.75 | 1.060 | positive |
| 2 | 55 | 1 | 64 | 160 | 77 | 270.0 | 1.99 | 0.003 | negative |
| 3 | 64 | 1 | 70 | 120 | 55 | 270.0 | 13.87 | 0.122 | positive |
| 4 | 55 | 1 | 64 | 112 | 65 | 300.0 | 1.08 | 0.003 | negative |
df['Result'] = df['Result'].map({'negative': 0, 'positive': 1})df['Result'].value_counts()1 810
0 509
Name: Result, dtype: int64EDA
- Gender
sns.countplot(x='Gender', hue='Result', data=df)
plt.title("Gender Distribution");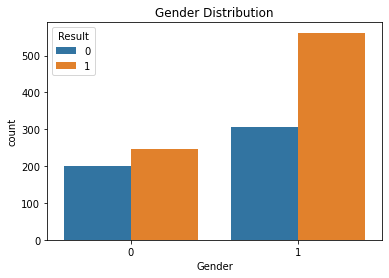
pd.crosstab(df['Gender'], df['Result'])| Result | 0 | 1 |
|---|---|---|
| Gender | ||
| 0 | 201 | 246 |
| 1 | 307 | 562 |
pd.crosstab(df['Gender'], df['Result'], normalize='index').round(2)| Result | 0 | 1 |
|---|---|---|
| Gender | ||
| 0 | 0.45 | 0.55 |
| 1 | 0.35 | 0.65 |
성별에 큰 의미가 없어보임.
Continuous Values
cols = ['Age', 'Heart rate','Systolic blood pressure','Diastolic blood pressure','Blood sugar','CK-MB','Troponin']
fig, axes = plt.subplots(3, 3, figsize=(16,10))
for i, col in enumerate(cols):
ax = axes[i//3, i%3]
sns.histplot(data=df, x=col, hue='Result', bins=20, ax=ax)
ax.set_title(col)
plt.tight_layout();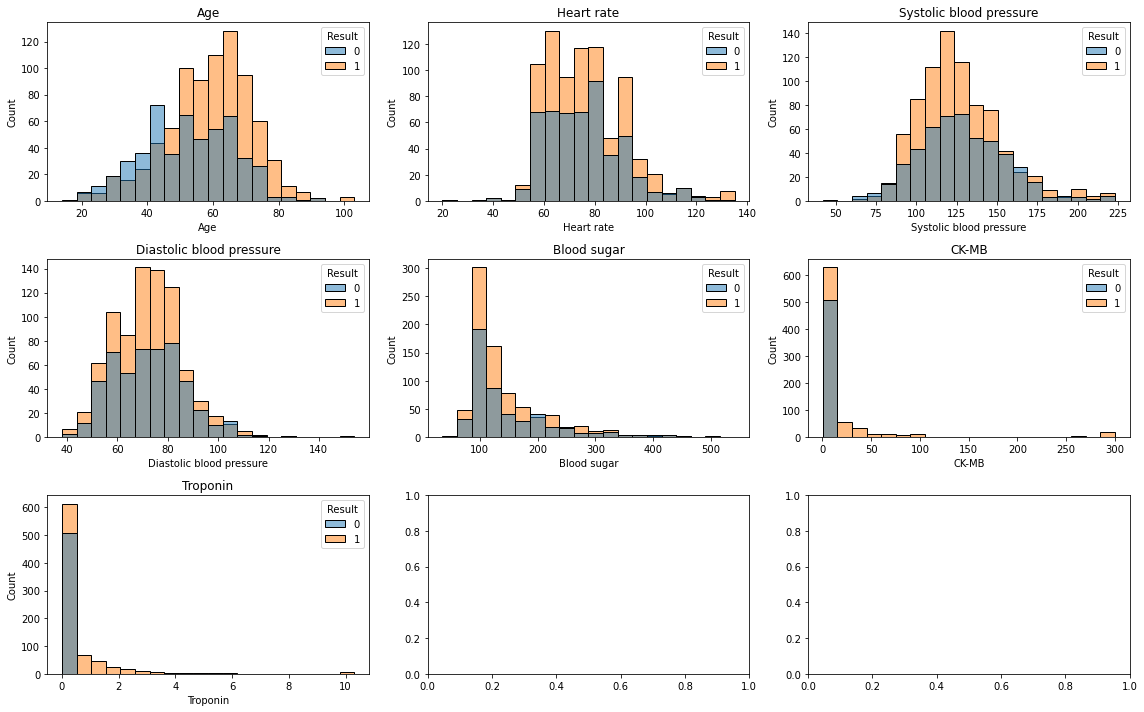
나이가 높을수록 심장 마비 영향이 높음
심박수는 영향이 없어보임
수축기혈압 systolic blood pressure은 영향 없어 보임
확장기혈압 diastolic blood pressure도 영향 없어 보임.
혈당도 영향 없어 보임
심장 근육속 존재하는 효소인 CK-MB도 영향 없어보임.
Troponin도..
Data distribution check
fig, axes = plt.subplots(3, 3, figsize=(16,12))
for i, col in enumerate(cols):
ax = axes[i//3, i%3]
sns.boxplot(x='Result', y=col, data=df, ax=ax)
ax.set_title(col)
plt.tight_layout();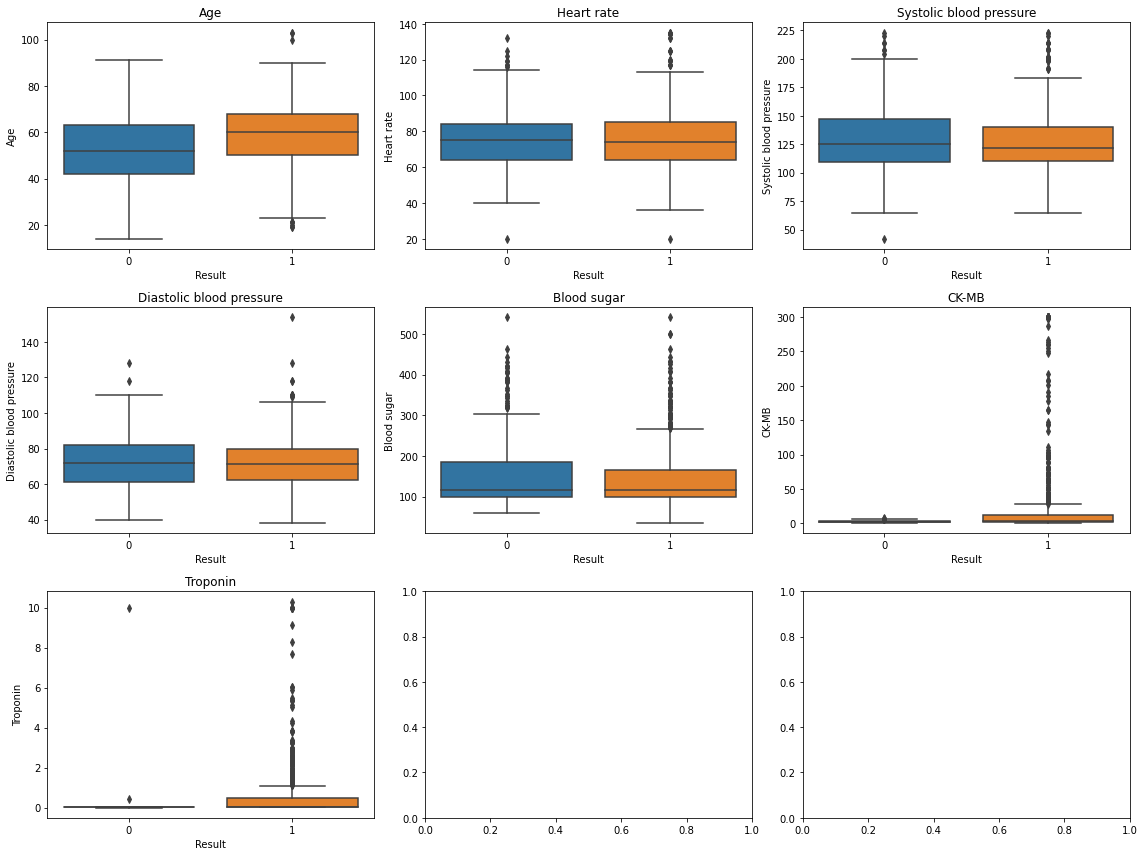
- 모두 심장마비 유무에 평균 차이가 없어보이나 이상치나 skewed된 모습이 보임
df.groupby('Result')[cols].mean().round(2)| Age | Heart rate | Systolic blood pressure | Diastolic blood pressure | Blood sugar | CK-MB | Troponin | |
|---|---|---|---|---|---|---|---|
| Result | |||||||
| 0 | 52.13 | 75.85 | 127.83 | 72.40 | 149.89 | 2.56 | 0.03 |
| 1 | 58.77 | 76.06 | 126.70 | 72.11 | 144.72 | 23.32 | 0.57 |
- ttest
for col in cols:
g0 = df[df['Result']==0][col]
g1 = df[df['Result']==1][col]
t,p = ttest_ind(g0,g1)
print(col, "p-value:", round(p,2))Heart rate p-value: 0.8
Systolic blood pressure p-value: 0.45
Diastolic blood pressure p-value: 0.73
Blood sugar p-value: 0.23
CK-MB p-value: 0.0
Troponin p-value: 0.0CK-MB, Troponin만 유의한 차이가 있음
data outliers
Values outside physiologically plausible ranges (e.g., heart rate > 250 bpm) were treated as data entry errors and set to missing.
df['Heart rate'].describe()count 1319.000000
mean 78.336619
std 51.630270
min 20.000000
25% 64.000000
50% 74.000000
75% 85.000000
max 1111.000000
Name: Heart rate, dtype: float64df.sort_values('Heart rate', ascending=False).head(5)| Age | Gender | Heart rate | Systolic blood pressure | Diastolic blood pressure | Blood sugar | CK-MB | Troponin | Result | |
|---|---|---|---|---|---|---|---|---|---|
| 1069 | 32 | 0 | 1111 | 141 | 95 | 82.0 | 2.66 | 0.008 | 0 |
| 717 | 70 | 0 | 1111 | 141 | 95 | 138.0 | 3.87 | 0.028 | 1 |
| 63 | 45 | 1 | 1111 | 141 | 95 | 109.0 | 1.33 | 1.010 | 1 |
| 682 | 68 | 1 | 135 | 98 | 60 | 96.0 | 254.40 | 0.025 | 1 |
| 1012 | 65 | 1 | 135 | 98 | 60 | 162.0 | 7.67 | 0.025 | 1 |
df = df[df['Heart rate'] < 250]Logistic Regression
df['Troponin'].hist(bins=30)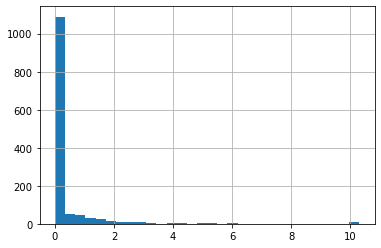
X = df[['Age','Gender','Heart rate',
'Systolic blood pressure',
'Diastolic blood pressure',
'Blood sugar','CK-MB','Troponin']]
y = df['Result']- train/test split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)- model
model = LogisticRegression(max_iter=1000)
model.fit(X_train, y_train)LogisticRegression(max_iter=1000)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
LogisticRegression(max_iter=1000)
- prediction
y_pred = model.predict(X_test)- score
print("Accuracy:", round(accuracy_score(y_test, y_pred),2))
y_prob = model.predict_proba(X_test)[:,1]
print("ROC AUC:", round(roc_auc_score(y_test, y_prob),2))Accuracy: 0.81
ROC AUC: 0.88coef = pd.DataFrame({
'Variable': X.columns,
'Coefficient': model.coef_[0],
'Odds Ratio': np.exp(model.coef_[0])
})print(round(coef,2)) Variable Coefficient Odds Ratio
0 Age 0.05 1.05
1 Gender 0.45 1.57
2 Heart rate 0.00 1.00
3 Systolic blood pressure -0.00 1.00
4 Diastolic blood pressure 0.00 1.00
5 Blood sugar -0.00 1.00
6 CK-MB 0.35 1.42
7 Troponin 6.84 932.77fpr, tpr, _ = roc_curve(y_test, y_prob)
plt.plot(fpr, tpr)
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.title("ROC Curve");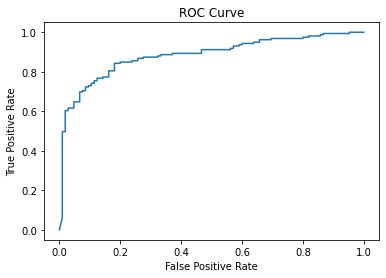
MLP, Multi-layer Perceptron
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)model = Sequential([
Dense(16, activation='relu', input_shape=(X_train.shape[1],)),
Dense(8, activation='relu'),
Dense(1, activation='sigmoid')
])model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])model.fit(X_train, y_train, epochs=50, batch_size=32, validation_split=0.2)Epoch 1/50
27/27 [==============================] - 0s 3ms/step - loss: 0.4792 - accuracy: 0.7539 - val_loss: 0.4921 - val_accuracy: 0.7488
Epoch 2/50
27/27 [==============================] - 0s 3ms/step - loss: 0.4714 - accuracy: 0.7693 - val_loss: 0.4865 - val_accuracy: 0.7488
Epoch 3/50
27/27 [==============================] - 0s 4ms/step - loss: 0.4630 - accuracy: 0.7741 - val_loss: 0.4784 - val_accuracy: 0.7583
Epoch 4/50
27/27 [==============================] - 0s 4ms/step - loss: 0.4552 - accuracy: 0.7836 - val_loss: 0.4770 - val_accuracy: 0.7488
Epoch 5/50
27/27 [==============================] - 0s 3ms/step - loss: 0.4469 - accuracy: 0.7895 - val_loss: 0.4674 - val_accuracy: 0.7536
Epoch 6/50
27/27 [==============================] - 0s 4ms/step - loss: 0.4395 - accuracy: 0.7848 - val_loss: 0.4623 - val_accuracy: 0.7488
Epoch 7/50
27/27 [==============================] - 0s 3ms/step - loss: 0.4332 - accuracy: 0.7919 - val_loss: 0.4594 - val_accuracy: 0.7630
Epoch 8/50
27/27 [==============================] - 0s 3ms/step - loss: 0.4276 - accuracy: 0.7943 - val_loss: 0.4543 - val_accuracy: 0.7583
Epoch 9/50
27/27 [==============================] - 0s 2ms/step - loss: 0.4217 - accuracy: 0.7967 - val_loss: 0.4465 - val_accuracy: 0.7630
Epoch 10/50
27/27 [==============================] - 0s 2ms/step - loss: 0.4161 - accuracy: 0.8026 - val_loss: 0.4445 - val_accuracy: 0.7725
Epoch 11/50
27/27 [==============================] - 0s 3ms/step - loss: 0.4113 - accuracy: 0.8050 - val_loss: 0.4387 - val_accuracy: 0.7678
Epoch 12/50
27/27 [==============================] - 0s 4ms/step - loss: 0.4065 - accuracy: 0.8026 - val_loss: 0.4341 - val_accuracy: 0.7773
Epoch 13/50
27/27 [==============================] - 0s 4ms/step - loss: 0.4014 - accuracy: 0.8050 - val_loss: 0.4296 - val_accuracy: 0.7773
Epoch 14/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3965 - accuracy: 0.8038 - val_loss: 0.4250 - val_accuracy: 0.7915
Epoch 15/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3930 - accuracy: 0.8062 - val_loss: 0.4217 - val_accuracy: 0.7820
Epoch 16/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3895 - accuracy: 0.8145 - val_loss: 0.4202 - val_accuracy: 0.7867
Epoch 17/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3858 - accuracy: 0.8062 - val_loss: 0.4139 - val_accuracy: 0.7867
Epoch 18/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3820 - accuracy: 0.8086 - val_loss: 0.4099 - val_accuracy: 0.7962
Epoch 19/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3788 - accuracy: 0.8169 - val_loss: 0.4096 - val_accuracy: 0.7773
Epoch 20/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3789 - accuracy: 0.8205 - val_loss: 0.4061 - val_accuracy: 0.7820
Epoch 21/50
27/27 [==============================] - 0s 2ms/step - loss: 0.3736 - accuracy: 0.8169 - val_loss: 0.4031 - val_accuracy: 0.8009
Epoch 22/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3698 - accuracy: 0.8181 - val_loss: 0.3977 - val_accuracy: 0.7962
Epoch 23/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3669 - accuracy: 0.8228 - val_loss: 0.3965 - val_accuracy: 0.8104
Epoch 24/50
27/27 [==============================] - 0s 2ms/step - loss: 0.3667 - accuracy: 0.8252 - val_loss: 0.3965 - val_accuracy: 0.7820
Epoch 25/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3617 - accuracy: 0.8264 - val_loss: 0.3904 - val_accuracy: 0.8104
Epoch 26/50
27/27 [==============================] - 0s 5ms/step - loss: 0.3594 - accuracy: 0.8264 - val_loss: 0.3893 - val_accuracy: 0.7962
Epoch 27/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3587 - accuracy: 0.8252 - val_loss: 0.3873 - val_accuracy: 0.8009
Epoch 28/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3560 - accuracy: 0.8276 - val_loss: 0.3859 - val_accuracy: 0.8104
Epoch 29/50
27/27 [==============================] - 0s 5ms/step - loss: 0.3534 - accuracy: 0.8288 - val_loss: 0.3848 - val_accuracy: 0.8104
Epoch 30/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3507 - accuracy: 0.8276 - val_loss: 0.3820 - val_accuracy: 0.8199
Epoch 31/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3483 - accuracy: 0.8252 - val_loss: 0.3795 - val_accuracy: 0.8152
Epoch 32/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3467 - accuracy: 0.8276 - val_loss: 0.3781 - val_accuracy: 0.8152
Epoch 33/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3445 - accuracy: 0.8371 - val_loss: 0.3755 - val_accuracy: 0.8199
Epoch 34/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3442 - accuracy: 0.8300 - val_loss: 0.3737 - val_accuracy: 0.8199
Epoch 35/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3414 - accuracy: 0.8312 - val_loss: 0.3730 - val_accuracy: 0.8152
Epoch 36/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3393 - accuracy: 0.8395 - val_loss: 0.3708 - val_accuracy: 0.8199
Epoch 37/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3385 - accuracy: 0.8359 - val_loss: 0.3701 - val_accuracy: 0.8199
Epoch 38/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3360 - accuracy: 0.8359 - val_loss: 0.3681 - val_accuracy: 0.8199
Epoch 39/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3341 - accuracy: 0.8371 - val_loss: 0.3670 - val_accuracy: 0.8246
Epoch 40/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3329 - accuracy: 0.8383 - val_loss: 0.3661 - val_accuracy: 0.8246
Epoch 41/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3311 - accuracy: 0.8371 - val_loss: 0.3639 - val_accuracy: 0.8152
Epoch 42/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3328 - accuracy: 0.8383 - val_loss: 0.3625 - val_accuracy: 0.8057
Epoch 43/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3291 - accuracy: 0.8419 - val_loss: 0.3623 - val_accuracy: 0.8152
Epoch 44/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3273 - accuracy: 0.8478 - val_loss: 0.3604 - val_accuracy: 0.8246
Epoch 45/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3270 - accuracy: 0.8395 - val_loss: 0.3588 - val_accuracy: 0.8104
Epoch 46/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3227 - accuracy: 0.8454 - val_loss: 0.3589 - val_accuracy: 0.8199
Epoch 47/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3243 - accuracy: 0.8454 - val_loss: 0.3551 - val_accuracy: 0.8246
Epoch 48/50
27/27 [==============================] - 0s 4ms/step - loss: 0.3216 - accuracy: 0.8442 - val_loss: 0.3557 - val_accuracy: 0.8104
Epoch 49/50
27/27 [==============================] - 0s 2ms/step - loss: 0.3185 - accuracy: 0.8490 - val_loss: 0.3563 - val_accuracy: 0.8152
Epoch 50/50
27/27 [==============================] - 0s 3ms/step - loss: 0.3178 - accuracy: 0.8430 - val_loss: 0.3511 - val_accuracy: 0.8341<keras.callbacks.History at 0x7fcde80caf70>y_pred = model.predict(X_test)9/9 [==============================] - 0s 891us/stepy_pred_class = (y_pred > 0.5).astype(int)print('accuracy: ',round(accuracy_score(y_test, y_pred_class),2))accuracy: 0.78print('roc: ',round(roc_auc_score(y_test, y_pred),2))roc: 0.87